
|

|
1
|
|
2
|
|
3
|
Click Add.
|
|
4
|
Click
|
|
5
|
|
6
|
Click
|
|
1
|
|
2
|
|
3
|
|
1
|
|
2
|
|
1
|
|
2
|
|
3
|
Click
|
|
4
|
Browse to the model’s Application Libraries folder and double-click the file carbon_deposition_parameters.txt.
|
|
1
|
In the Model Builder window, under Component 1 (comp1) right-click Definitions and choose Variables.
|
|
2
|
|
3
|
Click
|
|
4
|
Browse to the model’s Application Libraries folder and double-click the file carbon_deposition_variables.txt.
|
|
1
|
In the Model Builder window, under Component 1 (comp1) > Reaction Engineering (re) click 1: CH4 => C + 2 H2.
|
|
2
|
|
3
|
From the list, choose User defined.
|
|
4
|
|
1
|
|
2
|
|
3
|
In the Volumetric species table, enter the following settings:
|
|
1
|
|
2
|
|
4
|
|
5
|
|
6
|
|
7
|
|
8
|
|
1
|
|
2
|
|
3
|
|
4
|
|
1
|
|
2
|
|
1
|
In the Model Builder window, under Study 1 > Solver Configurations click Solution 1 - Copy 1 (sol2).
|
|
2
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
1
|
|
2
|
In the Settings window for Global, click Replace Expression in the upper-right corner of the y-Axis Data section. From the menu, choose Component 1 (comp1) > Reaction Engineering > re.c_a - Concentration - mol/m³.
|
|
3
|
|
4
|
|
5
|
|
1
|
In the Model Builder window, under Component 1 (comp1) > Reaction Engineering (re) click Species: a.
|
|
2
|
|
3
|
Select the Keep concentration/activity constant checkbox.
|
|
1
|
In the Model Builder window, under Study 1 > Solver Configurations click Solution 1 - Copy 1 (sol3).
|
|
2
|
|
1
|
|
2
|
In the Settings window for 1D Plot Group, type Concentration Comparison (re) in the Label text field.
|
|
3
|
|
4
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
1
|
|
2
|
|
3
|
|
4
|
Locate the Coloring and Style section. Find the Line style subsection. From the Line list, choose Dashed.
|
|
5
|
|
6
|
Locate the Legends section. In the table, enter the following settings:
|
|
7
|
|
1
|
In the Model Builder window, under Component 1 (comp1) > Reaction Engineering (re) click Species: C.
|
|
2
|
|
3
|
Select the Keep concentration/activity constant checkbox.
|
|
1
|
|
2
|
|
3
|
|
4
|
Locate the Physics Interfaces section. Find the Chemical species transport subsection. From the list, choose Transport of Concentrated Species: New.
|
|
5
|
|
6
|
|
7
|
|
1
|
|
2
|
|
3
|
|
4
|
Click
|
|
5
|
Browse to the model’s Application Libraries folder and double-click the file carbon_deposition_2D_variables.txt.
|
|
1
|
|
2
|
|
3
|
|
4
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
Click
|
|
1
|
|
2
|
On the object r2, select Points 2 and 3 only.
|
|
3
|
|
4
|
|
5
|
Click
|
|
1
|
|
2
|
Click in the Graphics window and then press Ctrl+A to select all objects.
|
|
3
|
|
4
|
Clear the Keep interior boundaries checkbox.
|
|
5
|
Click
|
|
1
|
|
2
|
On the object uni1, select Points 6 and 7 only.
|
|
3
|
|
4
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
Click
|
|
7
|
|
1
|
|
2
|
Go to the Add Physics window.
|
|
3
|
|
4
|
Click the Add to Component 2 button in the window toolbar.
|
|
5
|
|
1
|
|
2
|
|
4
|
|
5
|
|
6
|
In the Dependent variables (1) table, enter the following settings:
|
|
1
|
In the Model Builder window, under Component 2 (comp2) > Porosity Change (dode) click Distributed ODE 1.
|
|
2
|
|
3
|
|
1
|
|
2
|
|
3
|
|
1
|
In the Model Builder window, expand the Component 2 (comp2) > Chemistry (chem) node, then click Species: CH4.
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
|
7
|
|
8
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
|
7
|
|
8
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
|
7
|
|
8
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
|
1
|
In the Model Builder window, under Component 2 (comp2) click Transport of Concentrated Species (tcs).
|
|
2
|
In the Settings window for Transport of Concentrated Species, locate the Transport Mechanisms section.
|
|
3
|
Select the Mass transfer in porous media checkbox.
|
|
1
|
In the Model Builder window, expand the Transport of Concentrated Species (tcs) node, then click Fluid 1.
|
|
2
|
|
1
|
|
3
|
|
4
|
Click
|
|
6
|
|
1
|
|
3
|
|
4
|
|
1
|
|
1
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
Locate the Diffusion section. In the table, enter the following settings:
|
|
1
|
|
2
|
|
3
|
|
1
|
In the Model Builder window, expand the Component 2 (comp2) > Heat Transfer in Porous Media (ht) > Porous Medium 1 node, then click Fluid 1.
|
|
2
|
|
3
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
Locate the Heat Conduction, Porous Matrix section. From the ks list, choose User defined. In the associated text field, type kt_cat.
|
|
6
|
Locate the Thermodynamics, Porous Matrix section. From the ρs list, choose User defined. In the associated text field, type rho_cat.
|
|
7
|
|
1
|
In the Model Builder window, under Component 2 (comp2) > Heat Transfer in Porous Media (ht) click Heat Source 1.
|
|
3
|
|
4
|
|
1
|
|
1
|
|
1
|
|
3
|
|
4
|
|
1
|
|
1
|
|
2
|
|
3
|
Select the Enable porous media domains checkbox.
|
|
1
|
|
1
|
|
2
|
|
3
|
|
4
|
|
1
|
|
3
|
|
4
|
From the list, choose Fully developed flow.
|
|
5
|
|
1
|
|
1
|
|
3
|
|
4
|
|
1
|
|
2
|
|
3
|
In the Solve for column of the table, under Component 2 (comp2), clear the checkboxes for Chemistry (chem), Transport of Concentrated Species (tcs), Heat Transfer in Porous Media (ht), and Porosity Change (dode).
|
|
1
|
|
2
|
|
3
|
|
4
|
Locate the Physics and Variables Selection section. In the Solve for column of the table, under Component 1 (comp1), clear the checkbox for Reaction Engineering (re).
|
|
5
|
|
1
|
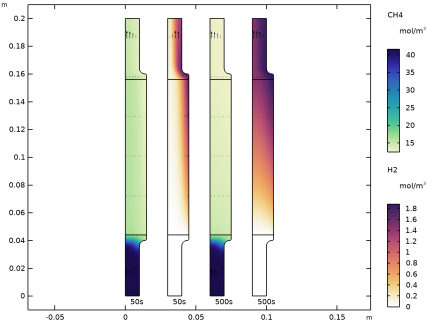
In the Model Builder window, under Results, Ctrl-click to select Concentration, CH4 (tcs), Concentration, CH4, 3D (tcs), Concentration, H2 (tcs), and Concentration, H2, 3D (tcs).
|
|
2
|
Right-click and choose Delete.
|
|
1
|
|
2
|
Go to the Result Templates window.
|
|
3
|
In the tree, select Study 2/Solution 4 (sol4) > Transport of Concentrated Species > Plot array: Concentrations, CH4, H2 (tcs).
|
|
4
|
Click the Add Result Template button in the window toolbar.
|
|
1
|
|
2
|
|
3
|
|
4
|
|
1
|
In the Model Builder window, expand the Plot array: Concentrations, CH4, H2 (tcs) node, then click CH4.
|
|
2
|
|
3
|
|
1
|
|
2
|
|
3
|
|
1
|
|
2
|
|
3
|
|
1
|
In the Model Builder window, under Results > Plot array: Concentrations, CH4, H2 (tcs), Ctrl-click to select Surface, CH4, Total Flux, CH4, CH4, Surface, H2, Total Flux, H2, and H2.
|
|
2
|
Right-click and choose Duplicate.
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
1
|
|
2
|
|
3
|
|
4
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
|
7
|
|
1
|
|
2
|
|
1
|
|
2
|
|
1
|
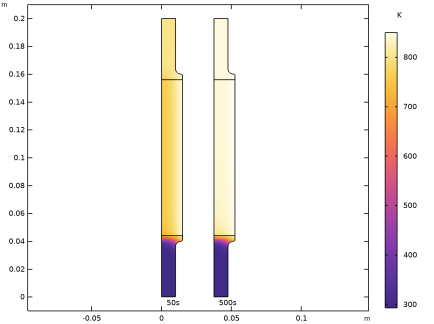
In the Model Builder window, under Results > Temperature (ht), Ctrl-click to select Surface 1 and 50s.
|
|
2
|
Right-click and choose Duplicate.
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
|
1
|
|
2
|
|
3
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
6
|
|
7
|
|
1
|
|
2
|
|
3
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
1
|
|
2
|
|
3
|
|
4
|
|
1
|
|
2
|
|
3
|
|
4
|
|
1
|
|
2
|
In the Settings window for 1D Plot Group, type Concentration CH4, Porous Catalyst Bed Center in the Label text field.
|
|
3
|
|
4
|
|
5
|
|
6
|
|
7
|
Locate the Plot Settings section.
|
|
8
|
Select the x-axis label checkbox. In the associated text field, type Length porous catalytic bed (m).
|
|
9
|
Select the y-axis label checkbox. In the associated text field, type Concentration (mol/m<sup>3</sup>).
|
|
10
|
|
1
|
|
2
|
In the Settings window for Line Graph, click Replace Expression in the upper-right corner of the y-Axis Data section. From the menu, choose Component 2 (comp2) > Transport of Concentrated Species > Species wCH4 > tcs.c_wCH4 - Molar concentration - mol/m³.
|
|
3
|
|
4
|
|
5
|
|
6
|
|
1
|
|
2
|
In the Settings window for Line Graph, click Replace Expression in the upper-right corner of the y-Axis Data section. From the menu, choose Component 2 (comp2) > Transport of Concentrated Species > Species wH2 > tcs.c_wH2 - Molar concentration - mol/m³.
|
|
3
|
Locate the Coloring and Style section. Find the Line style subsection. From the Line list, choose Dashed.
|
|
4
|
|
5
|
|
6
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
1
|
|
2
|
|
3
|
In the Solve for column of the table, under Component 2 (comp2), clear the checkbox for Porosity Change (dode).
|
|
1
|
|
2
|
|
3
|
|
4
|
|
1
|
|
2
|
|
3
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
1
|
|
2
|
|
3
|
|
1
|
In the Model Builder window, under Results > Reactor Overview right-click Surface 1 and choose Duplicate.
|
|
2
|
|
3
|
|
4
|
|
1
|
|
2
|
|
3
|
|
4
|
|
1
|
|
2
|
Click in the Graphics window and then press Ctrl+A to select all domains.
|
|
3
|
|
4
|
Clear the Evaluate the start cap checkbox.
|
|
5
|
Clear the Evaluate the end cap checkbox.
|
|
1
|
|
2
|
|
3
|
|
4
|
|
5
|
|
1
|
|
2
|
Right-click and choose Delete.
|